Scientific Background
Advances in technology favour development in medicine and biotechnology. One example is to support the discovery of new drugs thanks to the usage of the latest computer solutions.
One of the basic diagnostic tools is the analysis of changes in cell’s nuclei after drug treatment.
Due to the fact that the human body contains more than 30 trillion cells, which as themselves and the changes occurring inside them under the influence of the environment (including treatment), are a huge source of biological information [1], [2]. Moreover, the common problem in this field is the time needed to implement the product – circa 10 years. Hence, it is crucial to speed up the entire process [3], [4].
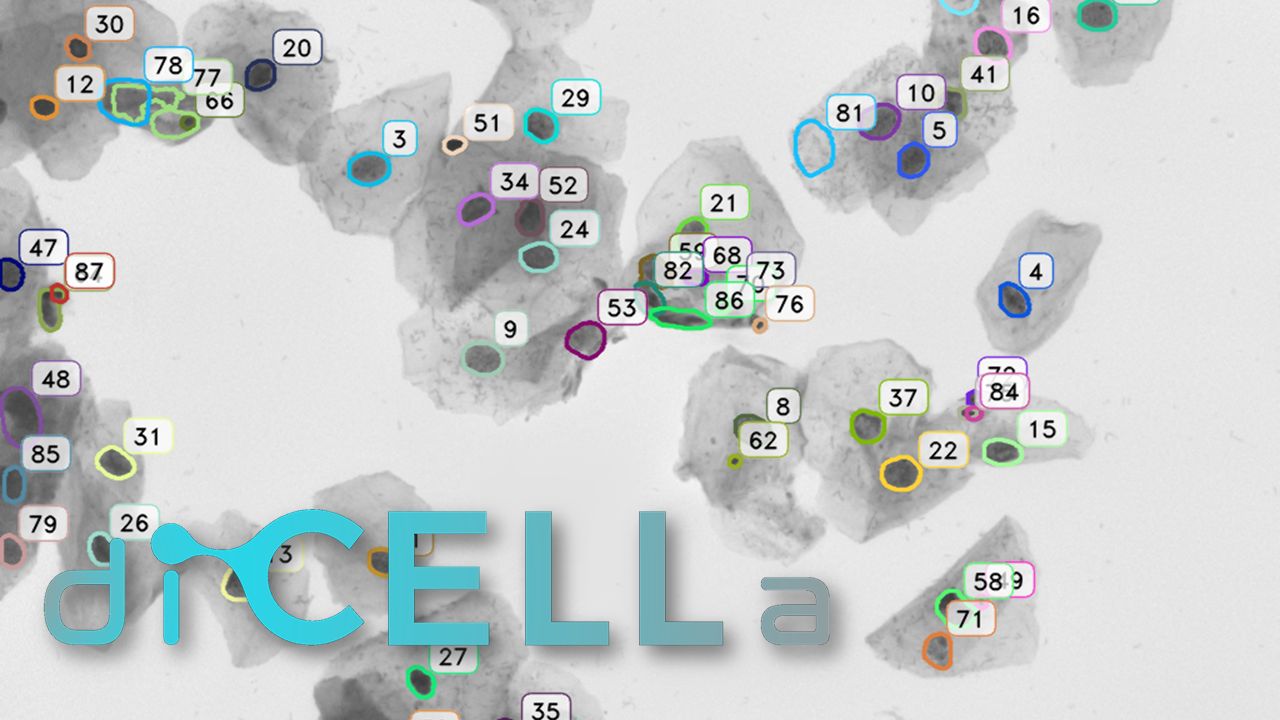
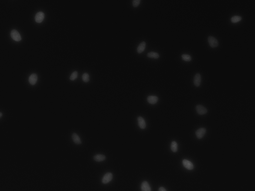
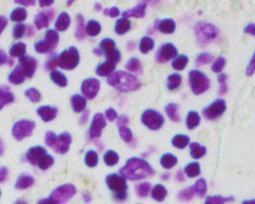
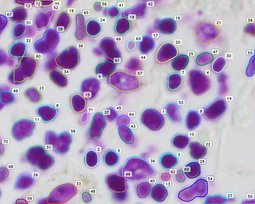
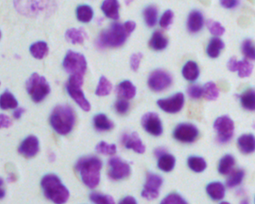
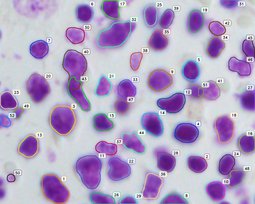
For this reason, we have developed an esemble model that allows to perform the automatic nuclei detection on the images, including fluorescent likewise steaming from a biopsy after staining by the most frequently used hematoxylin and eosin. We have chosen modern computer vision methods and deep learning to solve problems in this field of microscopy image analysis.
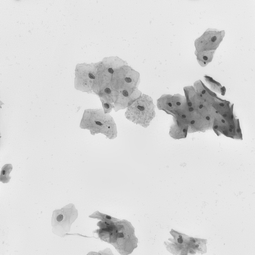
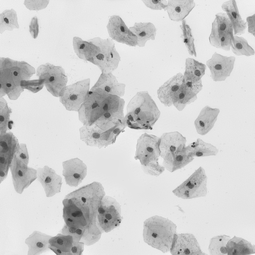
Undoubtedly, the vital examination in cancer diagnostic is biopsy analysis, which is becoming analytically useful after staining. The histopathologist work is difficult, time-consuming and tedious. Exact segmentation requires expert level knowledge and images may involve up to tens of thousands of nuclei needed to be labeled manually, so this solution could serve as the support system and which facilitate this type of task. It allows determining the clinically significant features, which are important in the diagnosis of cancer or inflammation, based on marked cell’s nuclei [5].
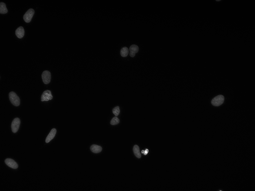
Our algorithm is dedicated to segmenting cell’s nuclei on the images acquired by the optical, phase-contrast and fluorescent microscope. It is robust to different contrast level or lighting [6]. Additionally, the fluorescence microscopy images are gaining in importance, because of the usefulness in molecular biology. They could be suitable for rapid high-throughput screening. The first stage in this research is the cell nuclei marking, which is the base for the future steps.
References:
- [1] Zhang Y., Ramaswami A., Bartolome C.: „DeepCell: Automating cell nuclei detection with.”, Stadford University, 2018
- [2] Alom, M. Z., Yakopcic, C., Taha, T. M., & Asari, V. K. (2018). Microscopic Nuclei Classification, Segmentation and Detection with improved Deep Convolutional Neural Network (DCNN) Approaches. arXiv preprint arXiv:1811.03447.
- [3] Alom, M. Z., Yakopcic, C., Taha, T. M., & Asari, V. K. (2018, July). Nuclei Segmentation with Recurrent Residual Convolutional Neural Networks based U-Net (R2U-Net). In NAECON 2018-IEEE National Aerospace and Electronics Conference (pp. 228-233). IEEE.
- [4] Zhou, Z., Siddiquee, M. M. R., Tajbakhsh, N., & Liang, J. (2018). Unet++: A nested u-net architecture for medical image segmentation. In Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support (pp. 3-11). Springer, Cham.
- [5] Naylor, P., Lae, M., Reyal, F., & Walter, T. (2017, April). Nuclei segmentation in histopathology images using deep neural networks. In 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017) (pp. 933-936). IEEE.
- [6] Dewan, M. A. A., Ahmad, M. O., & Swamy, M. N. S. (2014). A method for automatic segmentation of nuclei in phase-contrast images based on intensity, convexity and texture. IEEE transactions on biomedical circuits and systems, 8(5), 716-728.
- [7] Arslan, S., Ersahin, T., Cetin-Atalay, R., & Gunduz-Demir, C. (2013). Attributed relational graphs for cell nucleus segmentation in fluorescence microscopy images. IEEE transactions on medical imaging, 32(6), 1121-1131.
- [8] Vuola, A. O., Akram, S. U., & Kannala, J. (2019). Mask-RCNN and U-net Ensembled for Nuclei Segmentation. arXiv preprint arXiv:1901.10170.
- [9] Vuola, A. O., Akram, S. U., & Kannala, J. (2019). Mask-RCNN and U-net Ensembled for Nuclei Segmentation. arXiv preprint arXiv:1901.10170.